Projects
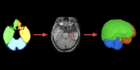
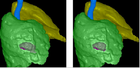
Image Analysis of Plant Panicles and Root Architecture

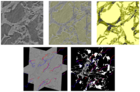
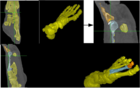
The morfology of plant organs, such as roots, leaves, and seeds, is closely related to relevant properties of the plant. For example, there are evidences that the architecture of a plant root system can indicate its tolerance to acid soils and its capacity to absorb nutrients, such as water and phosphorus. These properties are related to certain genes and the identification of those genes can improve the genetics of the plants. Genome-wide association studies (GWAS) look for the genes associated with a given trait of the plant. In this project, we are interested in phenotypic traits, which can be computed by 2D and 3D image analysis techniques. We are also interested in designing imaging systems suitable for phenotyping. We have investigated 3D image analysis of plant root systems (rice and sorghum) and 2D image analysis of rice panicles. The methods require image reconstruction, filtering, segmentation, and representation (e.g., skeletonization); phenotypic trait extraction; machine learning and pattern recognition techniques; and statistical data analysis. One important result of this project was PANorama --- a Linux-compatible, open-source software package for high-resolution panicle image acquisition, processing, and phenotyping. PANorama allowed the first study to assess inflorescence phenotypes of field-grown material.
Team:
Adán Echemendía MonteroAlexandre Xavier Falcão
Leon Kochian
Susan McCouch